
Software Tool for the Visualization of Transcriptional Modules
Hong Sun, Karen Lemmens, Tim Van den Bulcke, Kristof Engelen, Bart De Moor, Kathleen Marchal
Motivation:
Organisms are able to adapt their cellular machinery to changing environmental conditions. This complex cellular behavior is mediated by the underlying regulatory network. Previous studies have unveiled the modular and hierarchic organization of the transcriptional network. Module detection tools usually identify many overlapping modules. Indeed genes can be involved in multiple pathways. In addition, multiple pathways can be triggered in one particular environmental cue. Having a visual overview of how these modules overlap, gives insight in the structure of the biological system.The problem with visualizing overlapping modules simultaneously, however, is that the overlap in multiple dimensions complicates the choice of an appropriate layout. Therefore few tools exist that are capable of visualizing modules simultaneously.
Results:
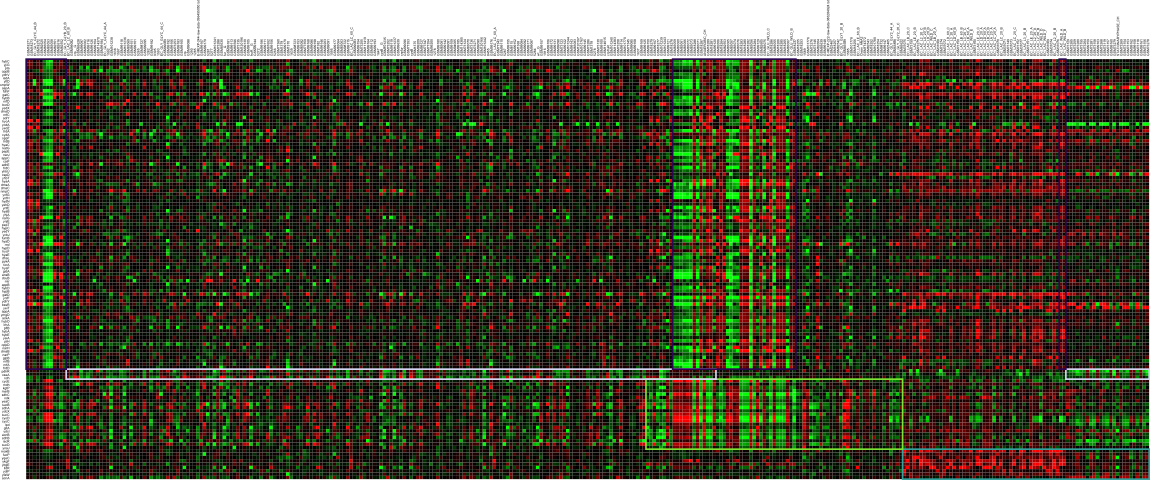
We have developed a tool, called ViTraM, that is capable of visualizing overlapping transcriptional modules in a very intuitive way. ViTraM allows for a dynamic visualization of overlapping transcriptional modules in a 2D gene-experiment matrix. Multiple methods are included for obtaining the optimal layout of the overlapping modules.
In addition to the previously developed tools for visualizing multiple modules, ViTraM also allows to display additional information on the regulatory program of the modules. The regulatory program consists of the transcription factors and their corresponding motifs. A first way of obtaining information on the regulatory program is by using the information from curated databases. This information can be used to further analyze modules inferred by biclustering algorithms. Secondly, information on the regulatory program can also be the outcome of a module inference tool itself. Both types of information on the regulatory program can be included by ViTraM. By visualizing not only the modules but also the regulatory program, ViTraM can provide more insight into the modules and makes the biological interpretation of the identified modules less complicated for biologists.
ViTraM requires as input an XML file containing information on all transcriptional modules. Expression data should also be loaded for some additional visualizations.
The example data files can be downloaded here.
In addition, we have developed software to create the required input files for ViTraM. The XMLCreator starts from the input and output data of DISTILLER and generates the required XML file and expression data. The XMLCreator software can be downloaded here, whereas the help file is available from here. Some example input and output data files can be found here.
The ViTraM 2.0 help file is available here. A Quick start guide can be downloaded here.
The current version of ViTraM is 2.0, released on Oct 1st, 2009.
The previous version of ViTraM is 1.0, released on Feb 15, 2009.
ViTraM 2.0 can be downloaded here.
ViTraM runs on both the Windows and Linux platform. Developed in JAVA, it is expected to work under other operating systems that support the JRE (1.5 or higher) and with sufficient memory depending on the size of the input data.
After downloading the package, please follow these steps:
1. Unzip the downloaded file
2. Open the unzipped folder
3. Depending on the OS:
>>Windows:
If you have a small data set, you can directly double click on the bat file vitram_Windows_512M.bat or vitram_Windows_2G.bat depending on the characteristic of your computer. Or file ViTraM.jar to run the software without assigning extra memory
Or open a command line window, go to the directory where you put ViTraM, and execute the command "java -jar -Xms256M -Xmx512M ViTraM.jar" in the folder in which the files are stored.
>>Linux:
Install X-Win32, VNC or other visualization software for Linux
Run ViTraM in a terminal with command "java -jar -Xms256m -Xmx2g ViTraM.jar".
Please allocate sufficient memory for ViTraM to run your data.
If you use ViTraM in your research, please cite the following publications:
H. Sun, K. Lemmens, T. Van den Buckle, K. Engelen, B. De Moor, K. Marchal. "ViTraM: Visualization of Transcriptional Modules" (2009). Bioinformatics, 25(18):2450-2451; doi:10.1093/bioinformatics/btp400.
H. Sun, K. Lemmens, T. Van den Buckle, K. Engelen, B. De Moor, K. Marchal. "Layout and Post-Processing of Transcriptional Modules". IEEE Computer Society, 10.1109/IJCBS.2009.95, 116-121 (2009). In Proceedings of the International Joint Conference on Bioinformatics, Systems Biology and Intelligent Computing.